单细胞代谢分析scMetabolism包
scMetabolism是一个用于量化单细胞分辨率代谢活性的R软件包。

scMetabolism:单细胞代谢活动量化工具指南
scMetabolism 是由复旦大学中山医院高强教授团队开发的一个 R 软件包,专门用于在单细胞分辨率下定量化代谢活性。
1. 环境准备与安装
V5结构,自己使用过程中,进行了一步改动。本文代码参考来源:生物信息学小白。
依赖包安装
在安装 scMetabolism 之前,需要先安装一系列依赖的生物信息学和绘图工具包:
# 常规依赖包
install.packages(c("devtools", "data.table", "wesanderson", "Seurat", "AUCell", "GSEABase", "GSVA", "ggplot2", "rsvd"))
# 指定版本的 VISION 核心依赖
devtools::install_github("YosefLab/VISION@v2.1.0")
主程序安装
devtools::install_github("wu-yc/scMetabolism")
2. 核心分析流程 (以 Seurat 为例)
数据加载
目前该工具主要支持人类 (Human) 的 scRNA-seq 数据。
library(scMetabolism)
library(ggplot2)
library(rsvd)
# 加载 Seurat 对象(示例数据:PBMC)
load(file = "pbmc_demo.rda")
代谢活动量化
推荐使用 sc.metabolism.Seurat 函数进行整合分析。
# 自定义sc.metabolism.SeuratV5函数
sc.metabolism.SeuratV5 <- function (obj, method = "VISION", imputation = F, ncores = 2,
metabolism.type = "KEGG") {
countexp <- GetAssayData(obj, layer = 'counts')
countexp <- data.frame(as.matrix(countexp))
signatures_KEGG_metab <- system.file("data", "KEGG_metabolism_nc.gmt",
package = "scMetabolism")
signatures_REACTOME_metab <- system.file("data", "REACTOME_metabolism.gmt",
package = "scMetabolism")
if (metabolism.type == "KEGG") {
gmtFile <- signatures_KEGG_metab
cat("Your choice is: KEGG\n")
}
if (metabolism.type == "REACTOME") {
gmtFile <- signatures_REACTOME_metab
cat("Your choice is: REACTOME\n")
}
if (imputation == F) {
countexp2 <- countexp
}
if (imputation == T) {
cat("Start imputation...\n")
cat("Citation: George C. Linderman, Jun Zhao, Yuval Kluger. Zero-preserving imputation of scRNA-seq data using low-rank approximation. bioRxiv. doi: https://doi.org/10.1101/397588 \n")
result.completed <- alra(as.matrix(countexp))
countexp2 <- result.completed[[3]]
row.names(countexp2) <- row.names(countexp)
}
cat("Start quantify the metabolism activity...\n")
if (method == "VISION") {
library(VISION)
n.umi <- colSums(countexp2)
scaled_counts <- t(t(countexp2)/n.umi) * median(n.umi)
vis <- Vision(scaled_counts, signatures = gmtFile)
options(mc.cores = ncores)
vis <- analyze(vis)
signature_exp <- data.frame(t(vis@SigScores))
}
if (method == "AUCell") {
library(AUCell)
library(GSEABase)
cells_rankings <- AUCell_buildRankings(as.matrix(countexp2),
nCores = ncores, plotStats = F)
geneSets <- getGmt(gmtFile)
cells_AUC <- AUCell_calcAUC(geneSets, cells_rankings)
signature_exp <- data.frame(getAUC(cells_AUC))
}
if (method == "ssGSEA") {
library(GSVA)
library(GSEABase)
geneSets <- getGmt(gmtFile)
gsva_es <- gsva(as.matrix(countexp2), geneSets, method = c("ssgsea"),
kcdf = c("Poisson"), parallel.sz = ncores)
signature_exp <- data.frame(gsva_es)
}
cat("\nPlease Cite: \nYingcheng Wu, Qiang Gao, et al. Cancer Discovery. 2021. \nhttps://pubmed.ncbi.nlm.nih.gov/34417225/ \n\n")
# 将代谢评分存储到 Seurat 对象中
obj@assays$METABOLISM$score <- signature_exp
obj
}
运行函数
res <- sc.metabolism.SeuratV5(obj = sce,
method = "AUCell",
imputation = F,
ncores = 20,
metabolism.type = "KEGG")
- ●obj是包含UMI计数矩阵的修拉对象。
- ●method: VISION是默认方法。VISION\AUCell \ssgsea \gsva
- ●imputation允许用户选择在新陈代谢评分前输入数据。
- ●ncores是并行计算线程的数量。
- ●metabolism.type 其中KEGG包含85条代谢通路,REACTOME包含82条代谢通路。
- ●结果提取:分析完成后,代谢评分矩阵存储在 obj@assaysMETABOLISMscore 中。
# 查看前 10 个通路名称,确认 "Glycolysis / Gluconeogenesis" 是否完全一致
head(rownames(res@assays$METABOLISM$score), 10)
3. 结果可视化
该工具提供了三种内置函数,用于展示特定代谢通路的活性分布。
可视化类型 | 函数名称 | 核心参数说明 |
|---|---|---|
降维图 (UMAP/tSNE) | DimPlot.metabolism | pathway: 目标通路名称;dimention.reduction.type: 降维方式 |
气泡图 (DotPlot) | DotPlot.metabolism | phenotype: 分组依据(如 celltype);norm: 归一化方向 (x/y) |
箱体图 (BoxPlot) | BoxPlot.metabolism | pathway: 可传入通路列表;ncol: 每行显示的图表数 |
# 1.定义DimPlot.metabolismV5函数
DimPlot.metabolismV5 <- function(obj, pathway, dimention.reduction.type = "umap",
dimention.reduction.run = TRUE, size = 1) {
cat("\nPlease Cite: \nYingcheng Wu, Qiang Gao, et al. Cancer Discovery. 2021. \nhttps://pubmed.ncbi.nlm.nih.gov/34417225/ \n\n")
if (dimention.reduction.type == "umap") {
if (dimention.reduction.run == TRUE)
obj <- Seurat::RunUMAP(obj, reduction = "pca", dims = 1:40)
umap.loc <- obj@reductions[["umap"]]@cell.embeddings
row.names(umap.loc) <- colnames(obj)
signature_exp <- obj@assays$METABOLISM$score
input.pathway <- pathway
signature_ggplot <- data.frame(umap.loc, t(signature_exp[input.pathway, ]))
library(wesanderson)
pal <- wes_palette("Zissou1", 100, type = "continuous")
library(ggplot2)
plot <- ggplot(data = signature_ggplot, aes(x = umap_1, y = umap_2,
color = signature_ggplot[, 3])) +
geom_point(size = size) +
scale_fill_gradientn(colours = pal) +
scale_color_gradientn(colours = pal) +
labs(color = input.pathway) +
xlab("UMAP 1") + ylab("UMAP 2") +
theme(aspect.ratio = 1) +
theme(panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
panel.background = element_blank(),
axis.line = element_line(colour = "black"))
}
if (dimention.reduction.type == "tsne") {
if (dimention.reduction.run == TRUE)
obj <- Seurat::RunTSNE(obj, reduction = "pca", dims = 1:40)
tsne.loc <- obj@reductions[["tsne"]]@cell.embeddings
row.names(tsne.loc) <- colnames(obj)
signature_exp <- obj@assays$METABOLISM$score
input.pathway <- pathway
signature_ggplot <- data.frame(tsne.loc, t(signature_exp[input.pathway, ]))
pal <- wes_palette("Zissou1", 100, type = "continuous")
library(ggplot2)
plot <- ggplot(data = signature_ggplot, aes(x = tSNE_1, y = tSNE_2,
color = signature_ggplot[, 3])) +
geom_point(size = size) +
scale_fill_gradientn(colours = pal) +
scale_color_gradientn(colours = pal) +
labs(color = input.pathway) +
xlab("tSNE 1") + ylab("tSNE 2") +
theme(aspect.ratio = 1) +
theme(panel.grid.major = element_blank(),
panel.grid.minor = element_blank(),
panel.background = element_blank(),
axis.line = element_line(colour = "black"))
}
return(plot)
}
DimPlot
DimPlot.metabolismV5(obj = res,
pathway = "Glycolysis / Gluconeogenesis",
dimention.reduction.type = "umap",
dimention.reduction.run = F, size = 1)
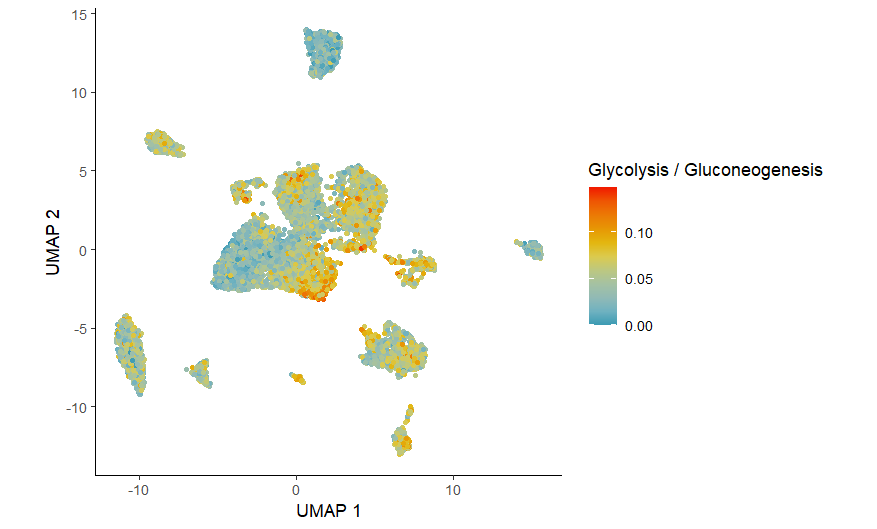
DotPlot
countexp.Seurat <- res
input.pathway <- rownames(countexp.Seurat@assays[["METABOLISM"]][["score"]])[1:9]
DotPlot.metabolism(obj = countexp.Seurat,
pathway = input.pathway,
phenotype = "celltype", # 使用meta.data中的celltype列
norm = "y") +
theme(axis.text.x = element_text(angle = 45, hjust = 1)) # 旋转x轴标签
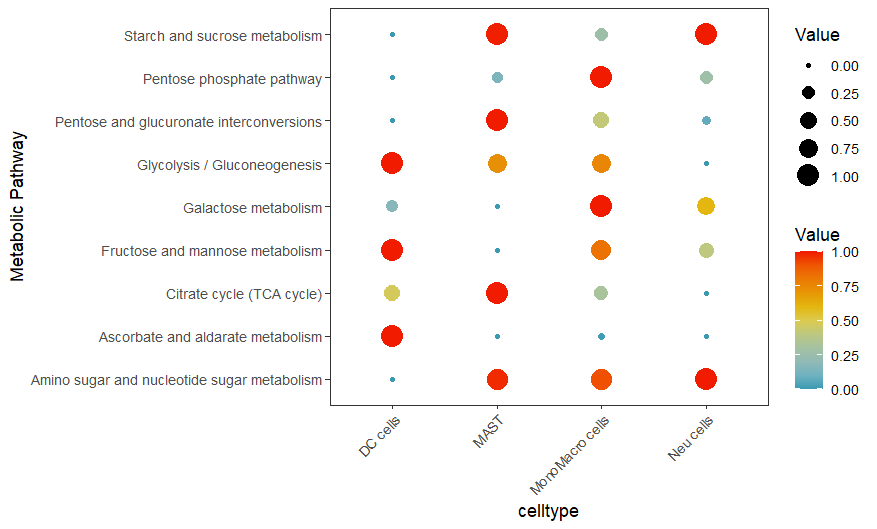
BoxPlot
BoxPlot.metabolism(obj = countexp.Seurat,
pathway = input.pathway,
phenotype = "celltype", #这个参数需按需修改
ncol = 3)
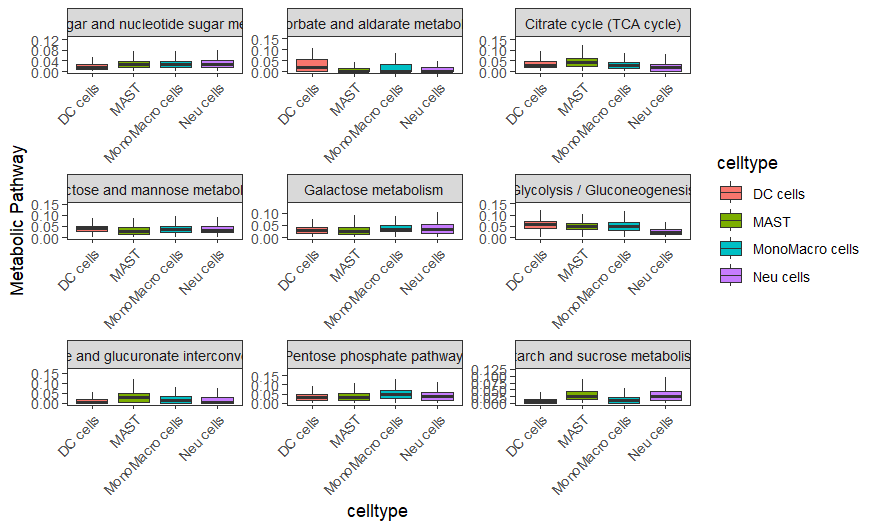
4. 其它信息
独立分析模式 (不推荐)
如果不使用 Seurat 对象,可以直接对计数矩阵 (Dataframe) 进行计算:
metabolism.matrix <- sc.metabolism(countexp = countexp, method = "AUCell", ...)
官网信息[1]
- ●核心算法文献:Wu et al., Cancer Discovery, 2021. (结直肠癌肝转移研究)
- ●在线版本:http://cancerdiversity.asia/scMetabolism/[2]
- ●联系方式:高强 (gaoqiang@fudan.edu.cn) 或 吴英成 (技术支持)
📚参考资料
[1]
官网信息: https://github.com/wu-yc/scMetabolism?tab=readme-ov-file
[2]
http://cancerdiversity.asia/scMetabolism/: http://cancerdiversity.asia/scMetabolism/
本文参与 腾讯云自媒体同步曝光计划,分享自微信公众号。
原始发表:2026-01-20,如有侵权请联系 cloudcommunity@tencent.com 删除
评论
登录后参与评论
推荐阅读
目录


