空转|CARD2-可基于基因集合进行spot注释,还可以提升到单细胞水平?
空转|CARD2-可基于基因集合进行spot注释,还可以提升到单细胞水平?

生信补给站
发布于 2023-08-25 10:35:46
发布于 2023-08-25 10:35:46
前面空转|CARD-结合scRNA解决空间转录组spot注释,还能增强空间精度?!介绍过了CARD 进行spot注释以及提升空间精度,本文介绍另外2个区别于其他注释软件的内容:
(1)reference-free version: CARDfree (基于基因集合进行空间spot注释)
(2)single cell resolution mapping (推断并绘制单细胞水平的空间spot注释)
空转相关数据使用CARD-1中使用的,先进行前面分析,然后接该推文的分析。
一 基因集合-spot注释
作者提供的CARDfree函数,可以不需要外部单细胞参考数据集,只需要用户输入之前已知的细胞类型标记的基因名称列表即可完成spot注释。
1,导入参考marker基因列表
这里使用CARD官网的markerList作为示例,可以根据自己的细胞类型需求进行自定义,保存为list。
load("./markerList.RData")
str(markerList)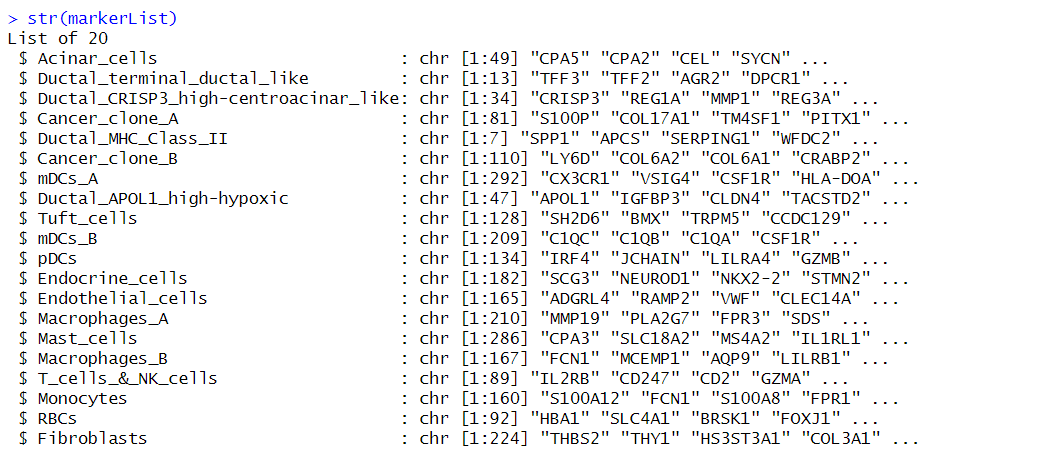
构建CARDfreeObject
CARDfree_obj = createCARDfreeObject(
markerList = markerList,
spatial_count = spatial_count,
spatial_location = spatial_location,
minCountGene = 100,
minCountSpot =5) ###创建CARDfree对象其中 空转数据的矩阵数据和位置信息分别存储在CARDfree_obj@spatial_countMat 和CARDfree_obj@spatial_location;marker列表存储在CARDfree_objj@markerList中。
2,CARDfree 解卷积
CARDfree_obj = CARD_refFree(CARDfree_obj)
colors = c("#FFD92F","#4DAF4A","#FCCDE5","#D9D9D9","#377EB8","#7FC97F","#BEAED4",
"#FDC086","#FFFF99","#386CB0","#F0027F","#BF5B17","#666666","#1B9E77",
"#D95F02","#7570B3","#E7298A","#66A61E","#E6AB02","#A6761D")
CARDfree_obj@Proportion_CARD = CARDfree_obj@Proportion_CARD[,c(8,10,14,2,1,6,12,18,7,13,20,19,16,17,11,15,4,9,3,5)]
colnames(CARDfree_obj@Proportion_CARD) = paste0("CT",1:20)
p1 <- CARD.visualize.pie(CARDfree_obj@Proportion_CARD,CARDfree_obj@spatial_location,colors = colors)
p1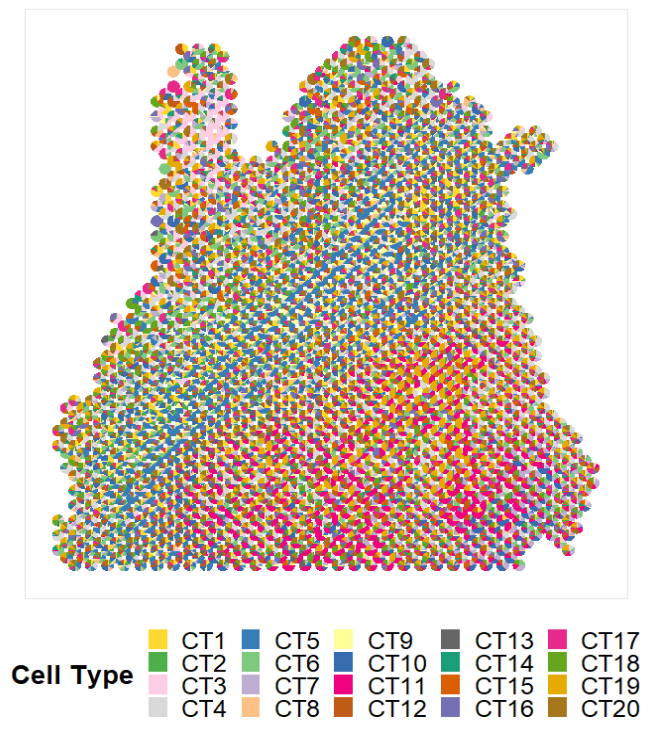
数据保存在CARDfree_obj@Proportion_CARD中。
二 单细胞分辨率空间图谱
CARD可以根据去卷积的参考单细胞转录组数据,从非单细胞分辨率空间转录组数据推断出每个空间spot位置的单细胞分辨率基因表达。
scMapping = CARD_SCMapping(CARD_obj,shapeSpot="Square",numCell=20,ncore=10)
print(scMapping)
#class: SingleCellExperiment
#dim: 20320 34200
#metadata(0):
#assays(1): counts
#rownames(20320): AL627309.1 AL627309.5 ... AC007325.4 AC007325.2
#rowData names(1): rownames(count_CT)
#colnames(34200): AAACAGGGTCTATATT-1:59x19:58.9649019359724x18.7704053990095
# GACGCATACCCGTCGG-1:59x19:59.2681436032053x18.9162972416363 ...
# GTCCCAATCATCCCGC-1:45x27:44.2986547542495x26.4444023868243
# TGCAGAACTATATCGT-1:45x27:44.3705854039158x26.3521195538112
#colData names(7): x y ... CT Cell
#reducedDimNames(0):
#mainExpName: NULL
#altExpNames(0):
### spatial location info and expression count of the single cell resolution data
library(SingleCellExperiment)
MapCellCords = as.data.frame(colData(scMapping))
count_SC = assays(scMapping)$counts
结果存储在singlecellexexperiment对象中;每个单细胞的空间位置信息以及它们与原始测量空间位置的关系存储在colData槽中。
注:至于推断的单细胞精度的空转注释 准确性如何,有待根据数据确定。
2,可视化
使用ggplot2进行可视化展示
df = MapCellCords
colors = c("#8DD3C7","#CFECBB","#F4F4B9","#CFCCCF","#D1A7B9","#E9D3DE","#F4867C","#C0979F",
"#D5CFD6","#86B1CD","#CEB28B","#EDBC63","#C59CC5","#C09CBF","#C2D567","#C9DAC3","#E1EBA0",
"#FFED6F","#CDD796","#F8CDDE")
p2 = ggplot(df, aes(x = x, y = y, colour = CT)) +
geom_point(size = 3.0) +
scale_colour_manual(values = colors) +
#facet_wrap(~Method,ncol = 2,nrow = 3) +
theme(plot.margin = margin(0.1, 0.1, 0.1, 0.1, "cm"),
panel.background = element_rect(colour = "white", fill="white"),
plot.background = element_rect(colour = "white", fill="white"),
legend.position="bottom",
panel.border = element_rect(colour = "grey89", fill=NA, size=0.5),
axis.text =element_blank(),
axis.ticks =element_blank(),
axis.title =element_blank(),
legend.title=element_text(size = 13,face="bold"),
legend.text=element_text(size = 12),
legend.key = element_rect(colour = "transparent", fill = "white"),
legend.key.size = unit(0.45, 'cm'),
strip.text = element_text(size = 15,face="bold"))+
guides(color=guide_legend(title="Cell Type"))
p2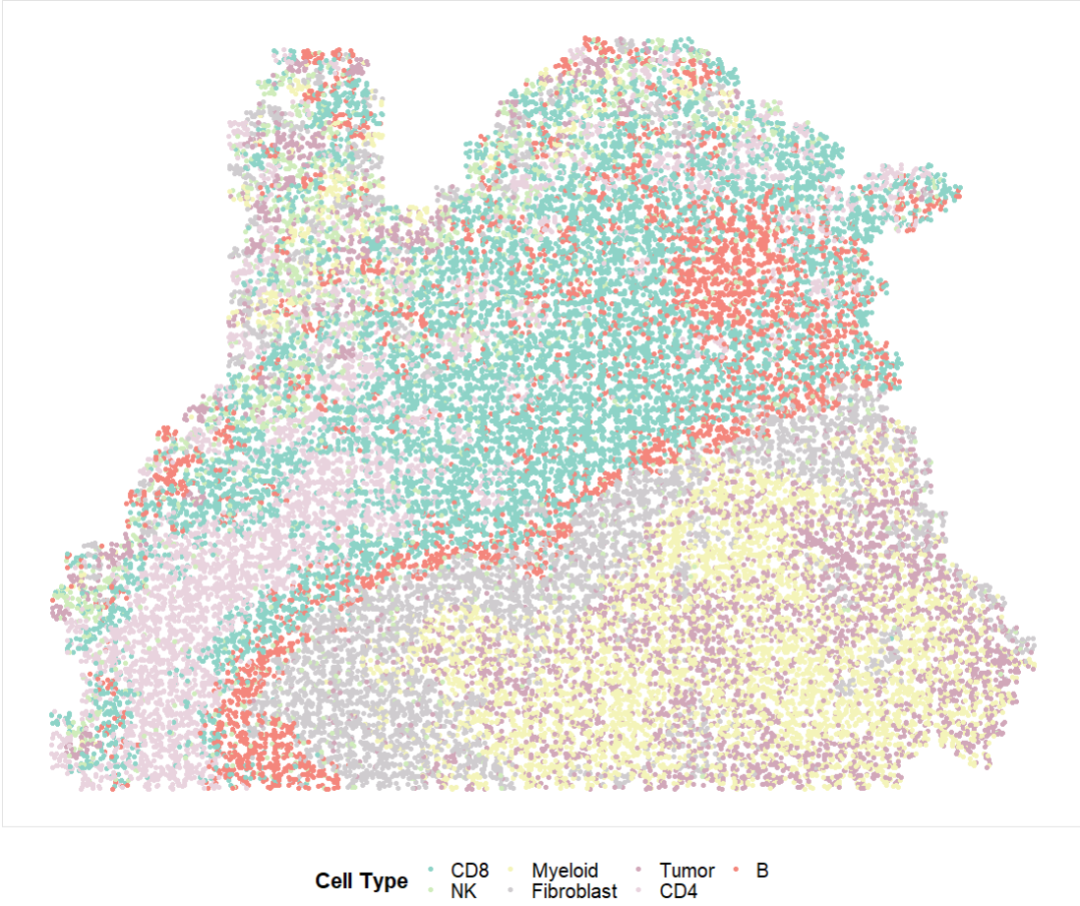
以上就完成了spot的空间注释,精度提升,使用基因列表进行spot注释 以及 单细胞精度的空转注释。
参考资料:
https://yingma0107.github.io/CARD/documentation/04_CARD_Example.html
本文参与 腾讯云自媒体同步曝光计划,分享自微信公众号。
原始发表:2023-08-22,如有侵权请联系 cloudcommunity@tencent.com 删除
评论
登录后参与评论
推荐阅读
目录

