🤗 Harmony | 超好用的单细胞测序数据合并(3'和5'数据合并)(二)
🤗 Harmony | 超好用的单细胞测序数据合并(3'和5'数据合并)(二)
生信漫卷
发布于 2023-02-24 13:51:14
发布于 2023-02-24 13:51:14
1写在前面
上一期我们介绍了常用的三种合并datasets的方法: 👇
Harmony;rliger;Seurat。
本期我们继续介绍其中的harmony包,如何用于3'和5'数据的合并。🤒
2用到的包
rm(list = ls())
library(Seurat)
library(SeuratDisk)
library(SeuratWrappers)
library(patchwork)
library(harmony)
library(rliger)
library(RColorBrewer)
library(tidyverse)
library(reshape2)
library(ggsci)
library(ggstatsplot)
3示例数据
这里我们提供1个3’ PBMC dataset和1个5’ PBMC dataset。🧐
matrix_3p <- Read10X_h5("./3p_pbmc10k_filt.h5",use.names = T)
matrix_5p <- Read10X_h5("./5p_pbmc10k_filt.h5",use.names = T)$`Gene Expression`
srat_3p <- CreateSeuratObject(matrix_3p,project = "pbmc10k_3p")
srat_5p <- CreateSeuratObject(matrix_5p,project = "pbmc10k_5p")
srat_3p
srat_5p
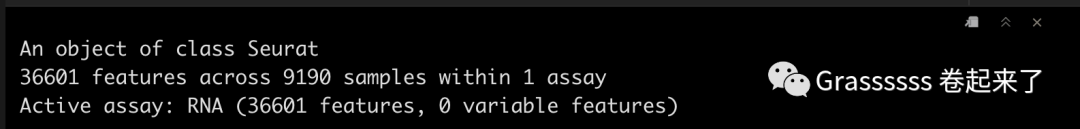
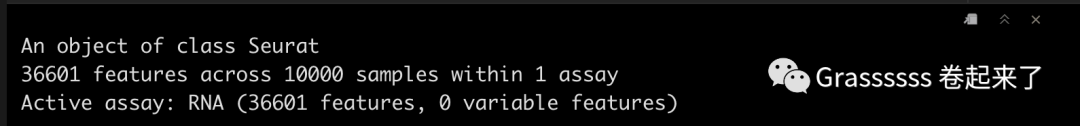
Note! 5' datset中还有一个assay,即VDJ data。🤜
4初步合并
4.1 简单合并
这里我们先用merge将2个数据集简单合并在一起。(这里我们默认做过初步过滤了哈,具体的大家可以看一下上期的教学。)😘
pbmc_harmony <- merge(srat_3p,srat_5p)
4.2 标准操作
我们在这里做一下Normalization,寻找高变基因等标准操作。👀
pbmc_harmony <- NormalizeData(pbmc_harmony, verbose = F)
pbmc_harmony <- FindVariableFeatures(pbmc_harmony, selection.method = "vst", nfeatures = 2000, verbose = F)
pbmc_harmony <- ScaleData(pbmc_harmony, verbose = F)
pbmc_harmony <- RunPCA(pbmc_harmony, npcs = 30, verbose = F)
pbmc_harmony <- RunUMAP(pbmc_harmony, reduction = "pca", dims = 1:30, verbose = F)
5harmony合并数据
5.1 合并前
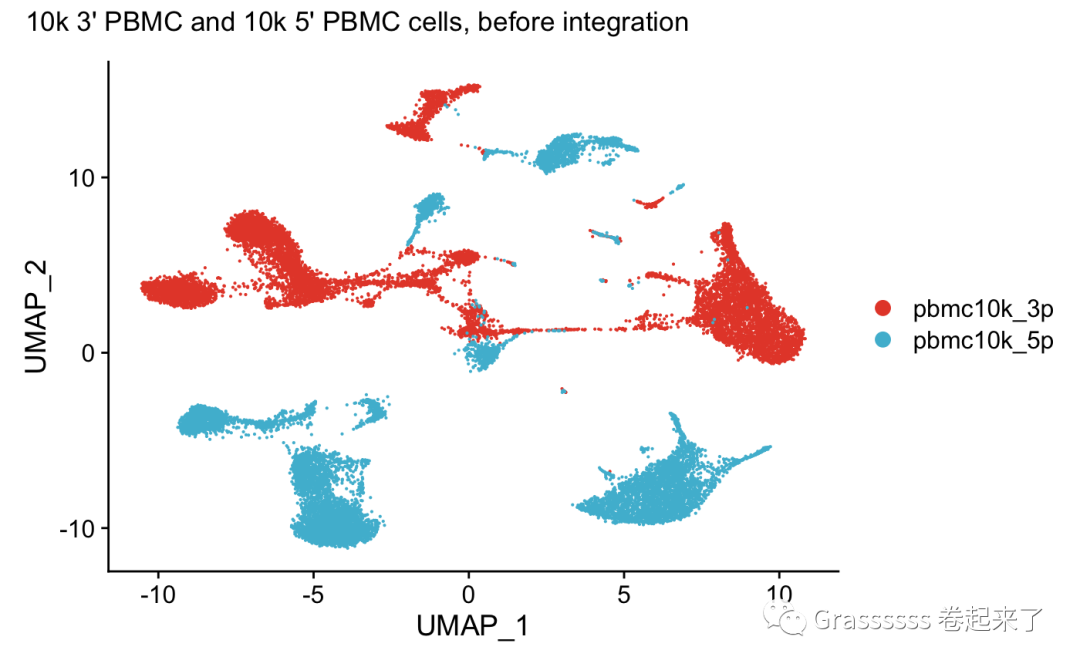
harmony合并前,PCA明显分离。🥲
DimPlot(pbmc_harmony, reduction = "umap") +
scale_color_npg()+
plot_annotation(title = "10k 3' PBMC and 10k 5' PBMC cells, before integration")
5.2 开始合并
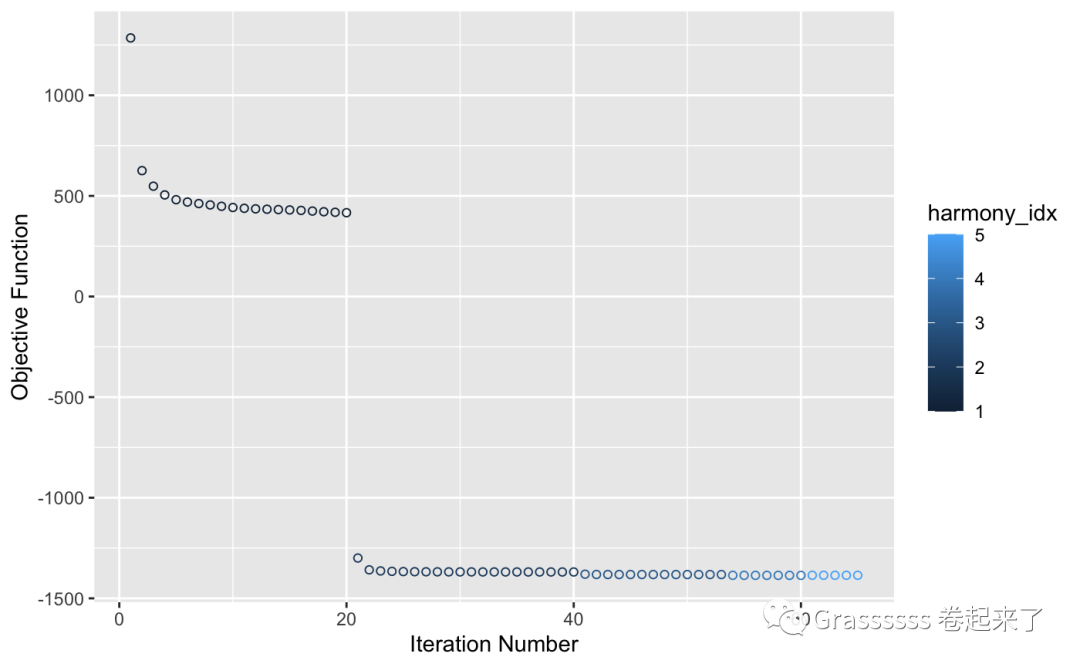
pbmc_harmony <- pbmc_harmony %>%
RunHarmony("orig.ident", plot_convergence = T)
5.3 查看信息
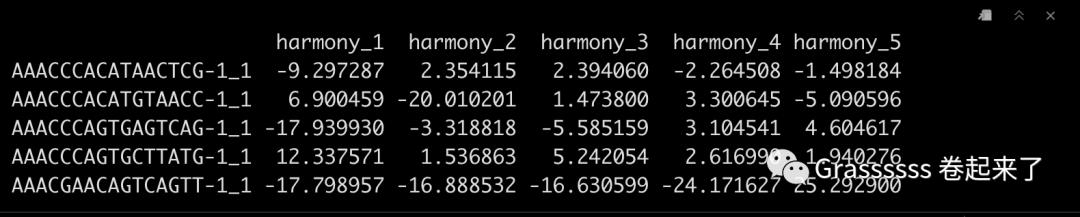
harmony_embeddings <- Embeddings(pbmc_harmony, 'harmony')
harmony_embeddings[1:5, 1:5]
5.4 可视化-harmony
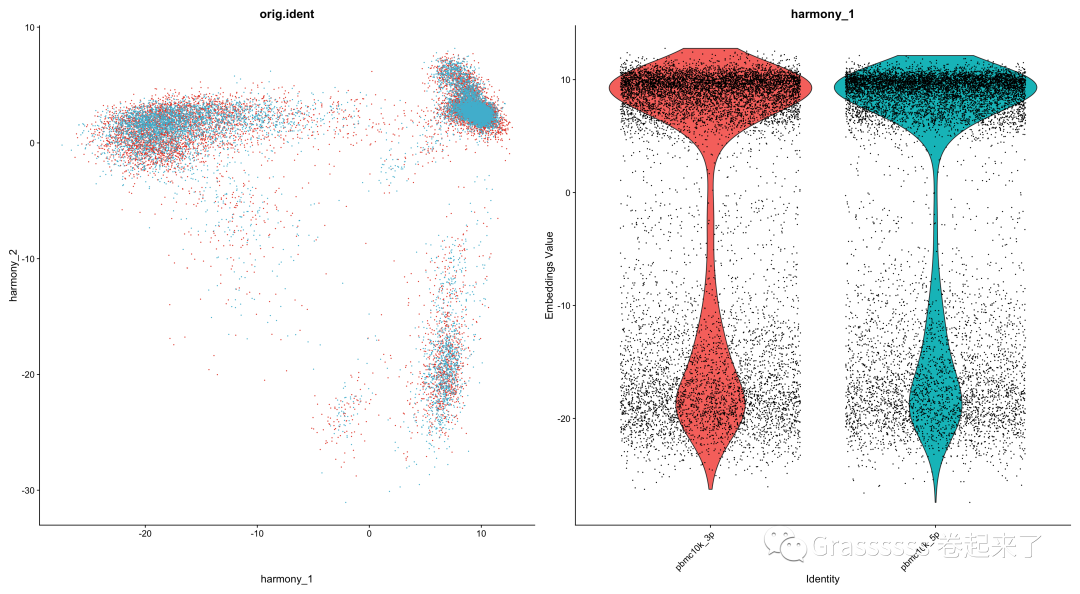
p1 <- DimPlot(object = pbmc_harmony, reduction = "harmony",
pt.size = .1, group.by = "orig.ident") +
scale_color_npg()+
NoLegend()
p2 <- VlnPlot(object = pbmc_harmony, features = "harmony_1",
group.by = "orig.ident", pt.size = .1) +
scale_color_npg()+
NoLegend()
p1 + p2
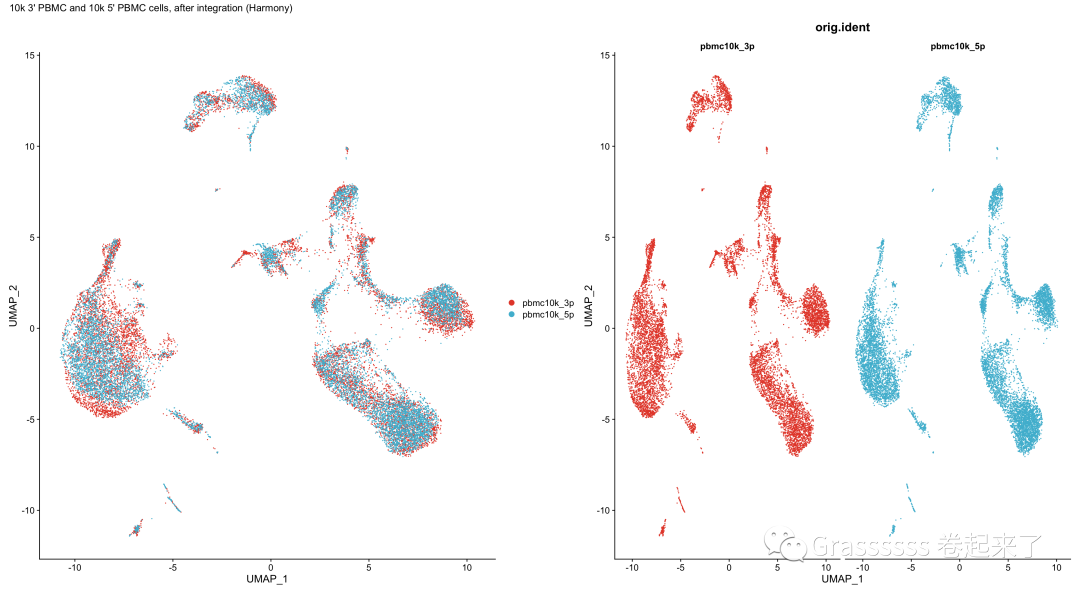
5.5 可视化-UMAP
harmony合并后,UMAP几乎重叠,但效果似乎没有Seurat包好。🤒
pbmc_harmony <- pbmc_harmony %>%
RunUMAP(reduction = "harmony", dims = 1:30, verbose = F)
pbmc_harmony <- SetIdent(pbmc_harmony,value = "orig.ident")
p1 <- DimPlot(pbmc_harmony,reduction = "umap") +
scale_color_npg()+
plot_annotation(title = "10k 3' PBMC and 10k 5' PBMC cells, after integration (Harmony)")
p2 <- DimPlot(pbmc_harmony, reduction = "umap",
group.by = "orig.ident", pt.size = .1,
split.by = 'orig.ident') +
scale_color_npg()+
NoLegend()
p1 + p2
6降维与聚类
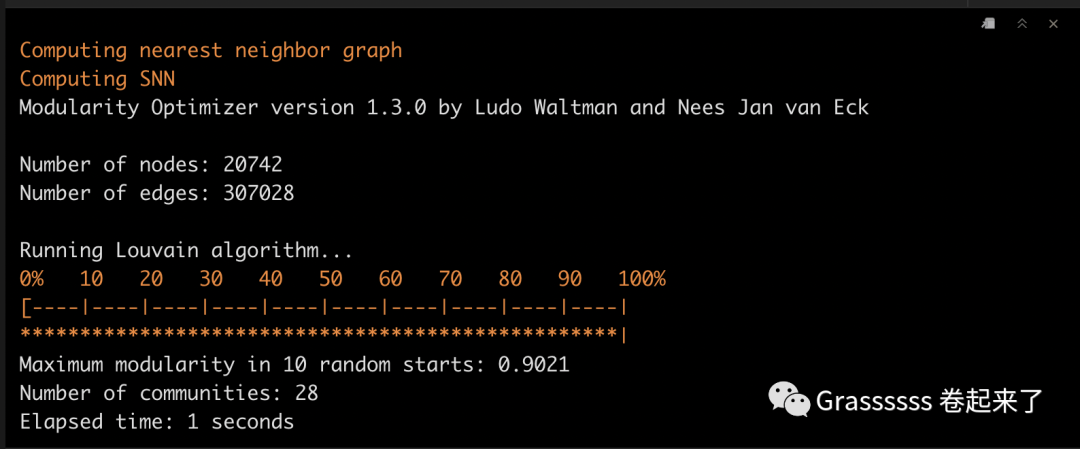
6.1 寻找clusters
pbmc_harmony <- pbmc_harmony %>%
FindNeighbors(reduction = "harmony", k.param = 10, dims = 1:30) %>%
FindClusters() %>%
identity()
6.2 聚类可视化
pbmc_harmony <- SetIdent(pbmc_harmony,value = "seurat_clusters")
ncluster <- length(unique(pbmc_harmony[[]]$seurat_clusters))
mycol <- colorRampPalette(brewer.pal(8, "Set2"))(ncluster)
DimPlot(pbmc_harmony,label = T,
cols = mycol, repel = T) +
NoLegend()
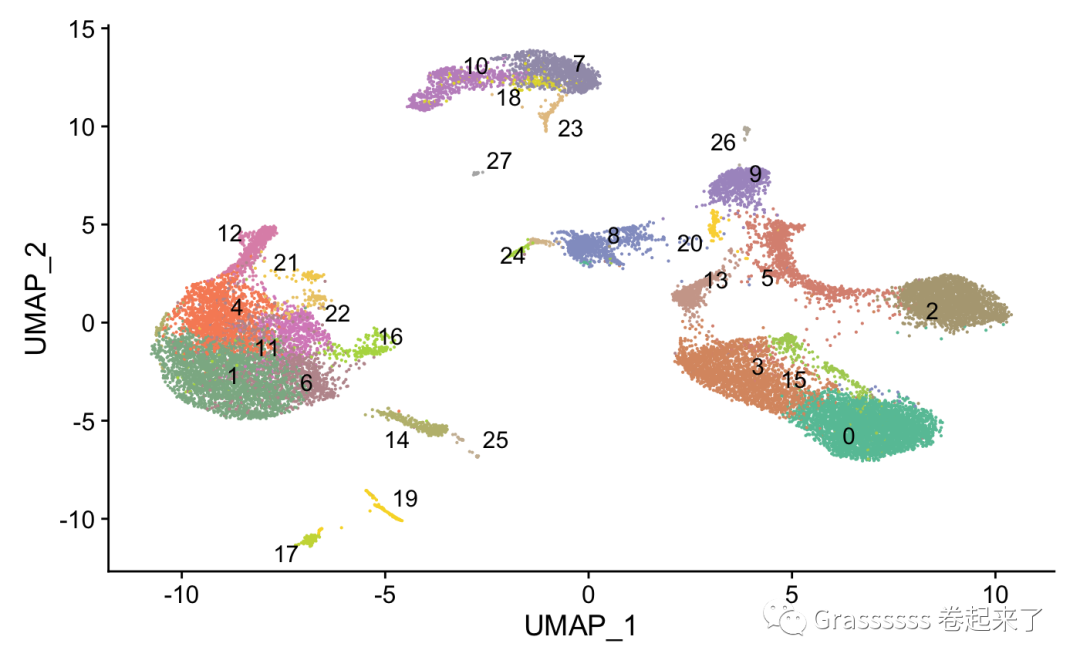
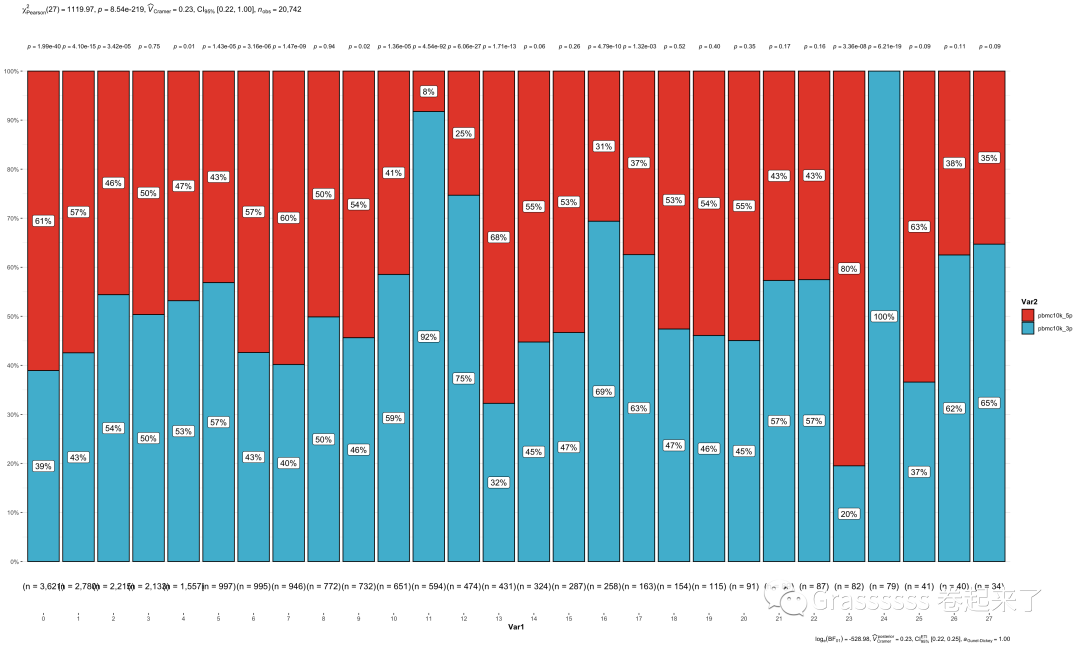
6.3 具体查看及可视化
我们看下各个clusters在两个datasets各有多少细胞。
count_table <- table(pbmc_harmony@meta.data$seurat_clusters, pbmc_harmony@meta.data$orig.ident)
count_table
#### 可视化
count_table %>%
as.data.frame() %>%
ggbarstats(x = Var2,
y = Var1,
counts = Freq)+
scale_fill_npg()
最后祝大家早日不卷!~
本文参与 腾讯云自媒体同步曝光计划,分享自微信公众号。
原始发表:2022-10-25,如有侵权请联系 cloudcommunity@tencent.com 删除
评论
登录后参与评论
推荐阅读
目录

